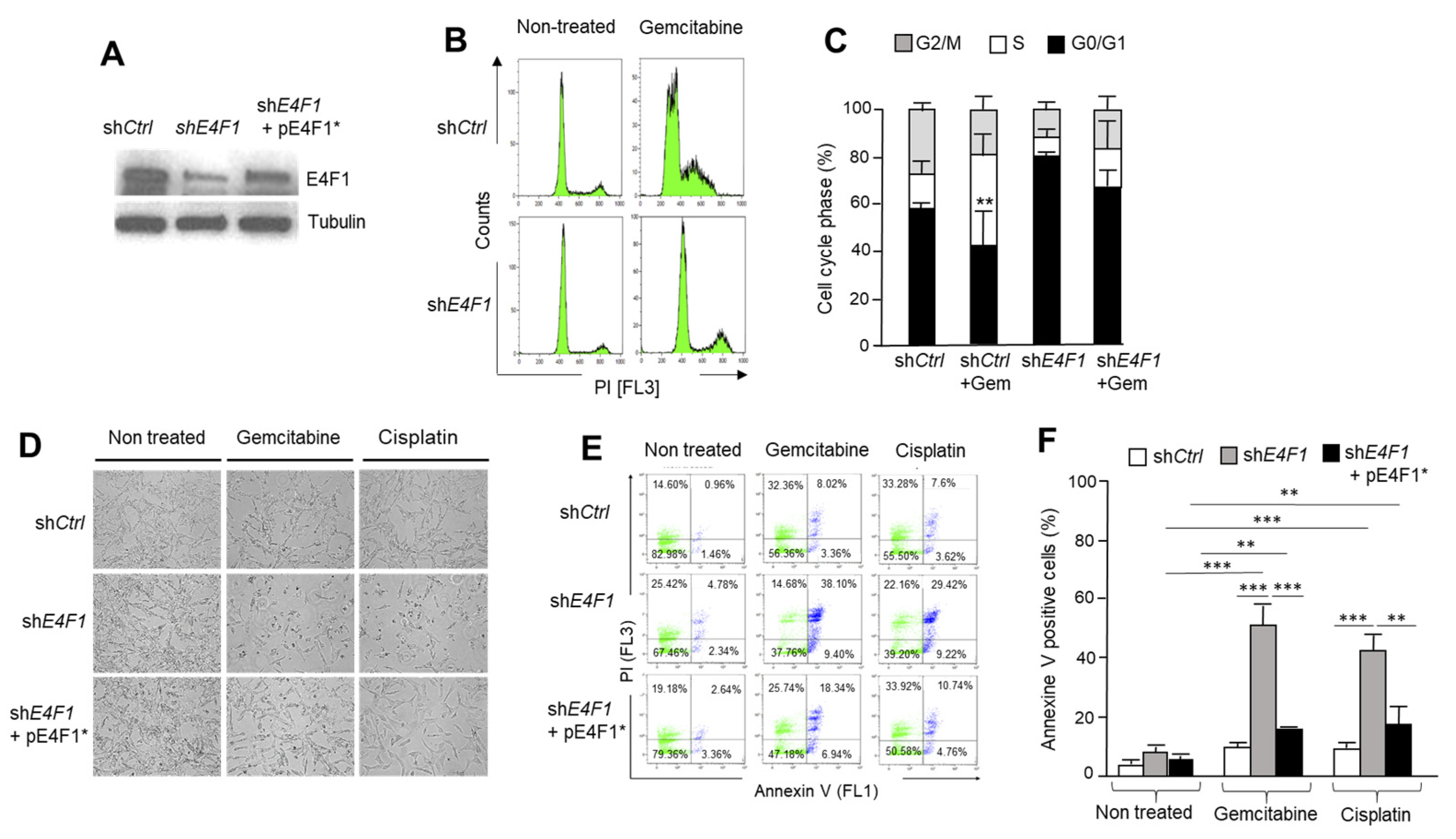
IJMS | Free Full-Text | Multi-Level Control of the ATM/ATR-CHK1 Axis by the Transcription Factor E4F1 in Triple-Negative Breast Cancer

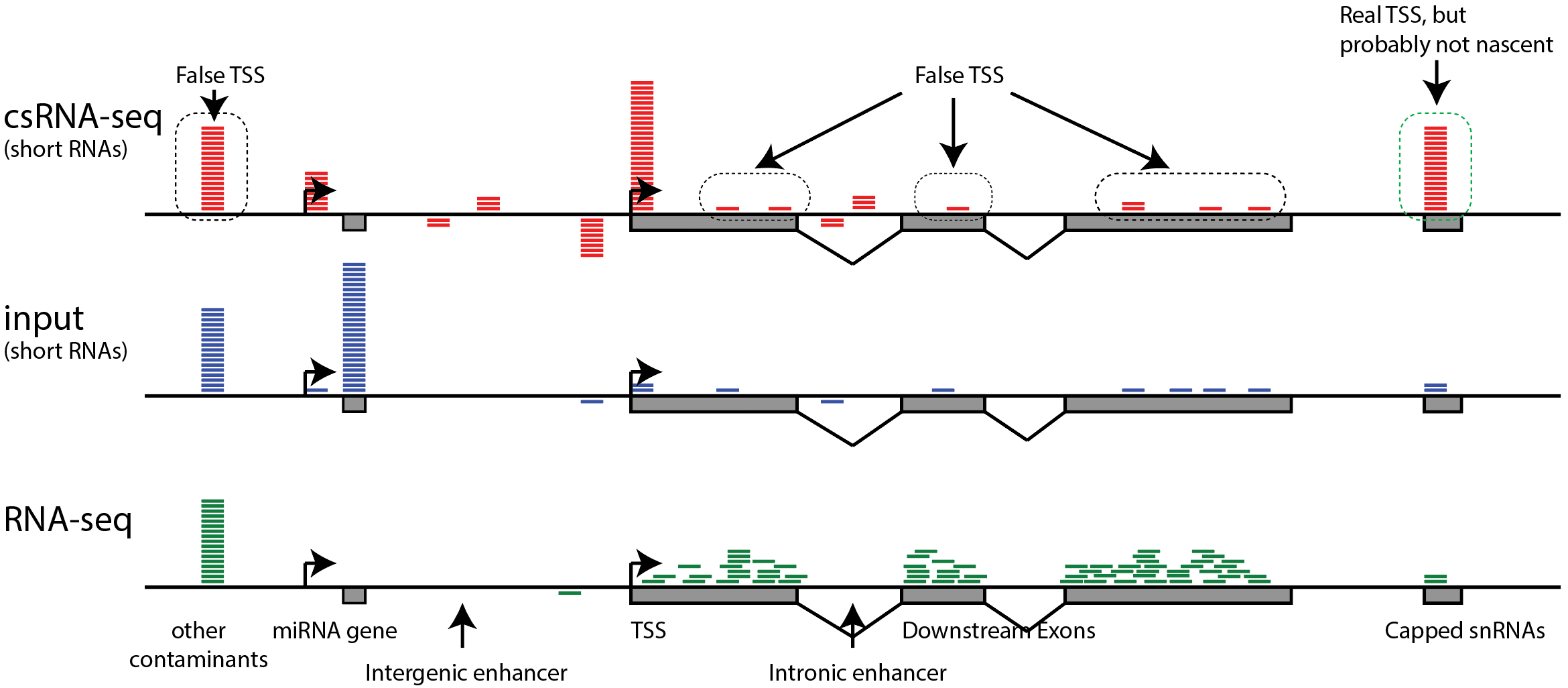
Frontiers | Mapping Mammalian Cell-type-specific Transcriptional Regulatory Networks Using KD-CAGE and ChIP-seq Data in the TC-YIK Cell Line
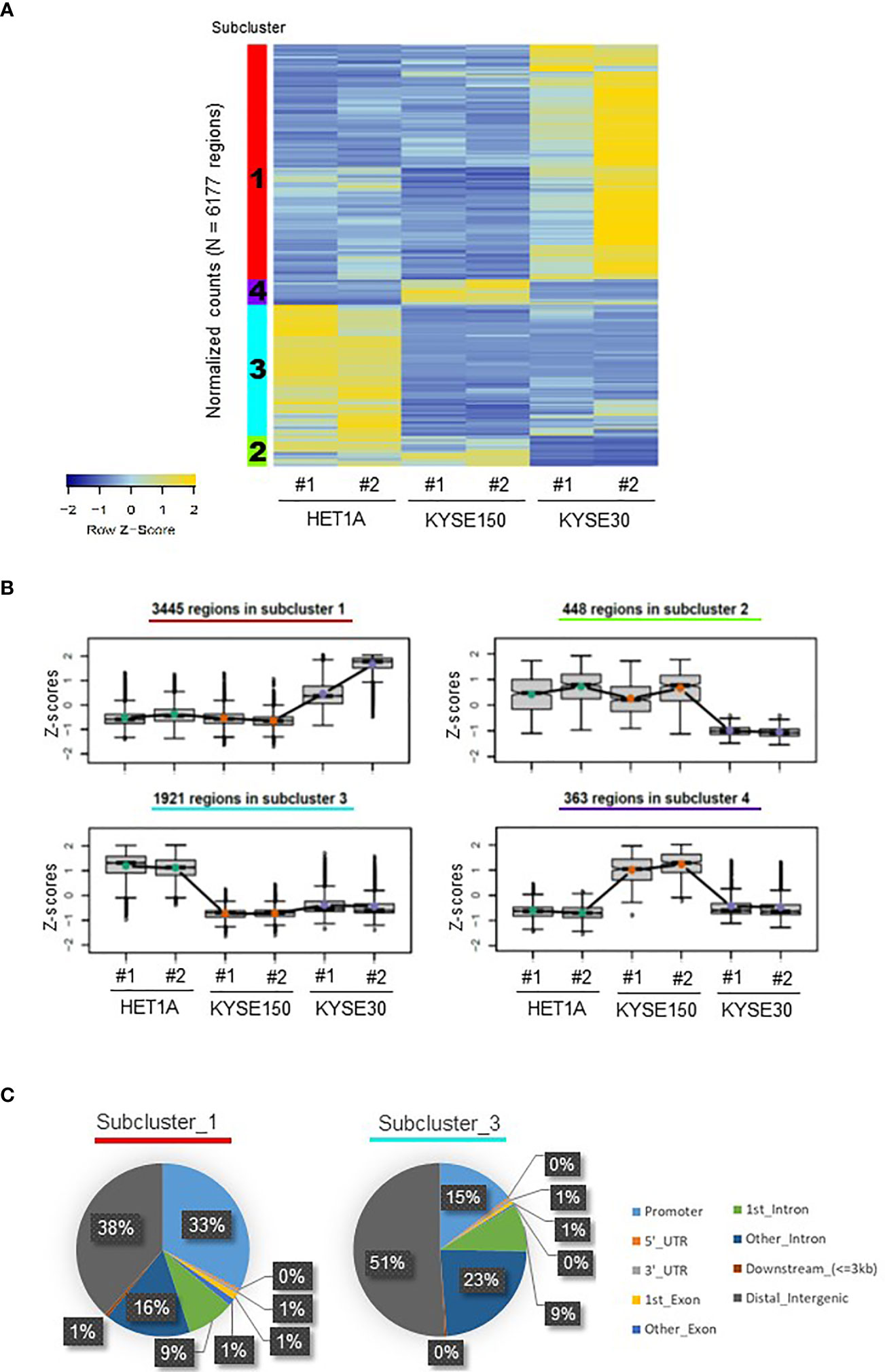
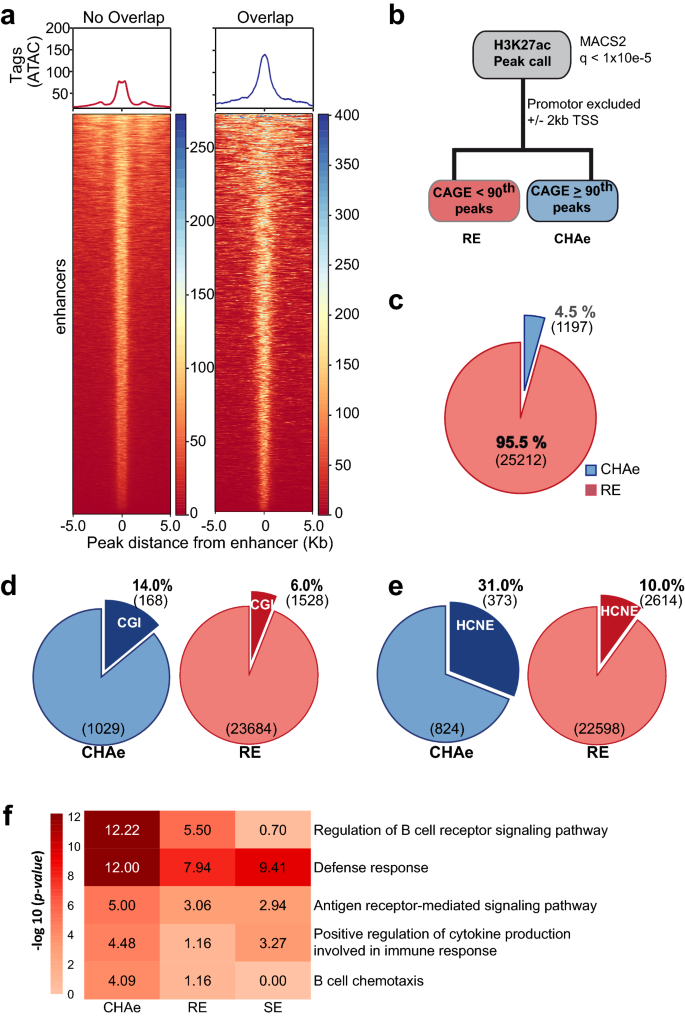
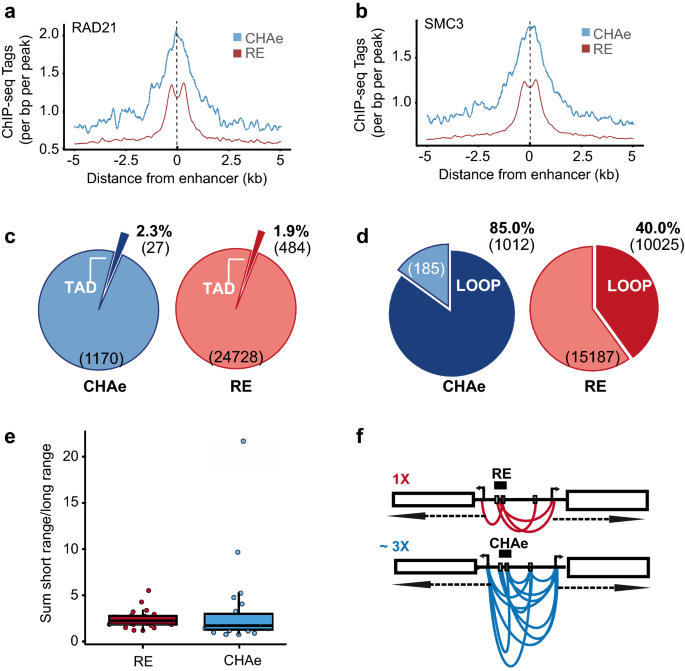
Frontiers | Identification of a Putative Enhancer RNA for EGFR in Hyper-Accessible Regions in Esophageal Squamous Cell Carcinoma Cells by Analysis of Chromatin Accessibility Landscapes
Alternative promoters in CpG depleted regions pervasively account for epigenetic misregulation of cancer transcriptomes
Mapping Mammalian Cell-type-specific Transcriptional Regulatory Networks Using KD-CAGE and ChIP-seq Data in the TC-YIK Cell Line
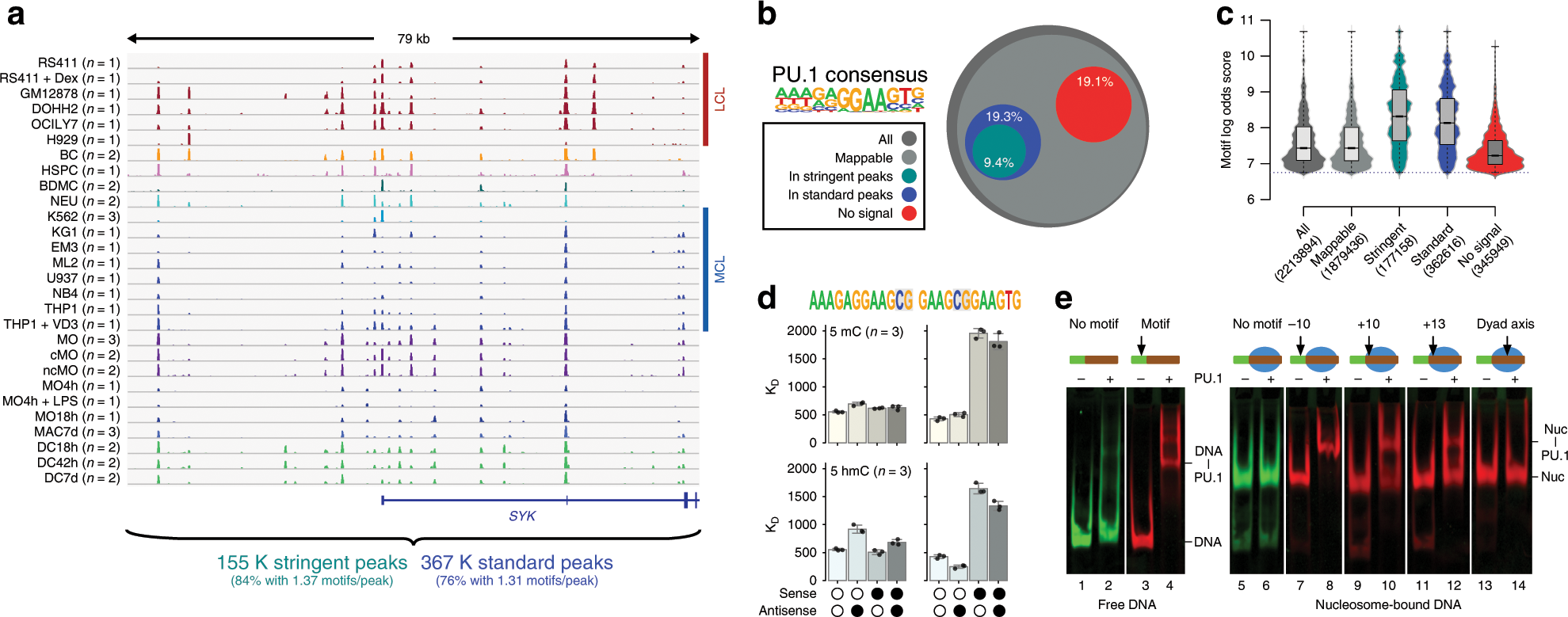
Mechanisms governing the pioneering and redistribution capabilities of the non-classical pioneer PU.1 | Nature Communications

Lactobacillus plantarum-derived indole-3-lactic acid ameliorates colorectal tumorigenesis via epigenetic regulation of CD8+ T cell immunity - ScienceDirect

Endogenous retroviruses co-opted into divergently transcribed regulatory elements shape the regulatory landscape of embryonic stem cells | bioRxiv

Endogenous retroviruses co-opted into divergently transcribed regulatory elements shape the regulatory landscape of embryonic stem cells | bioRxiv
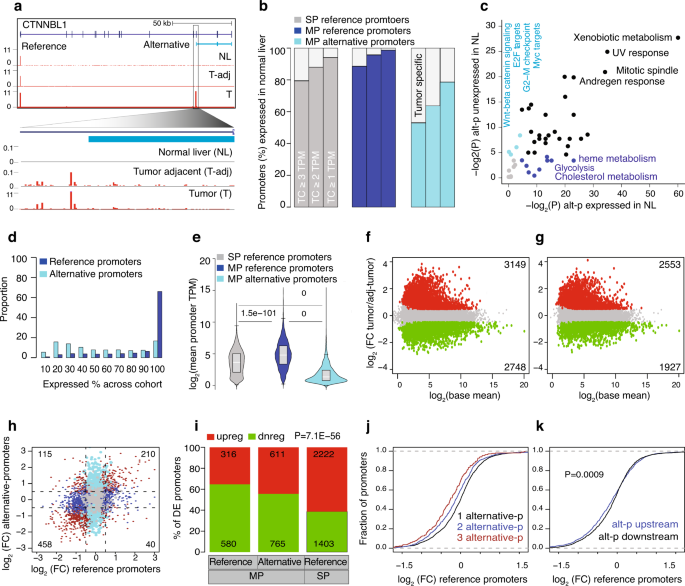
Alternative promoters in CpG depleted regions are prevalently associated with epigenetic misregulation of liver cancer transcriptomes | Nature Communications
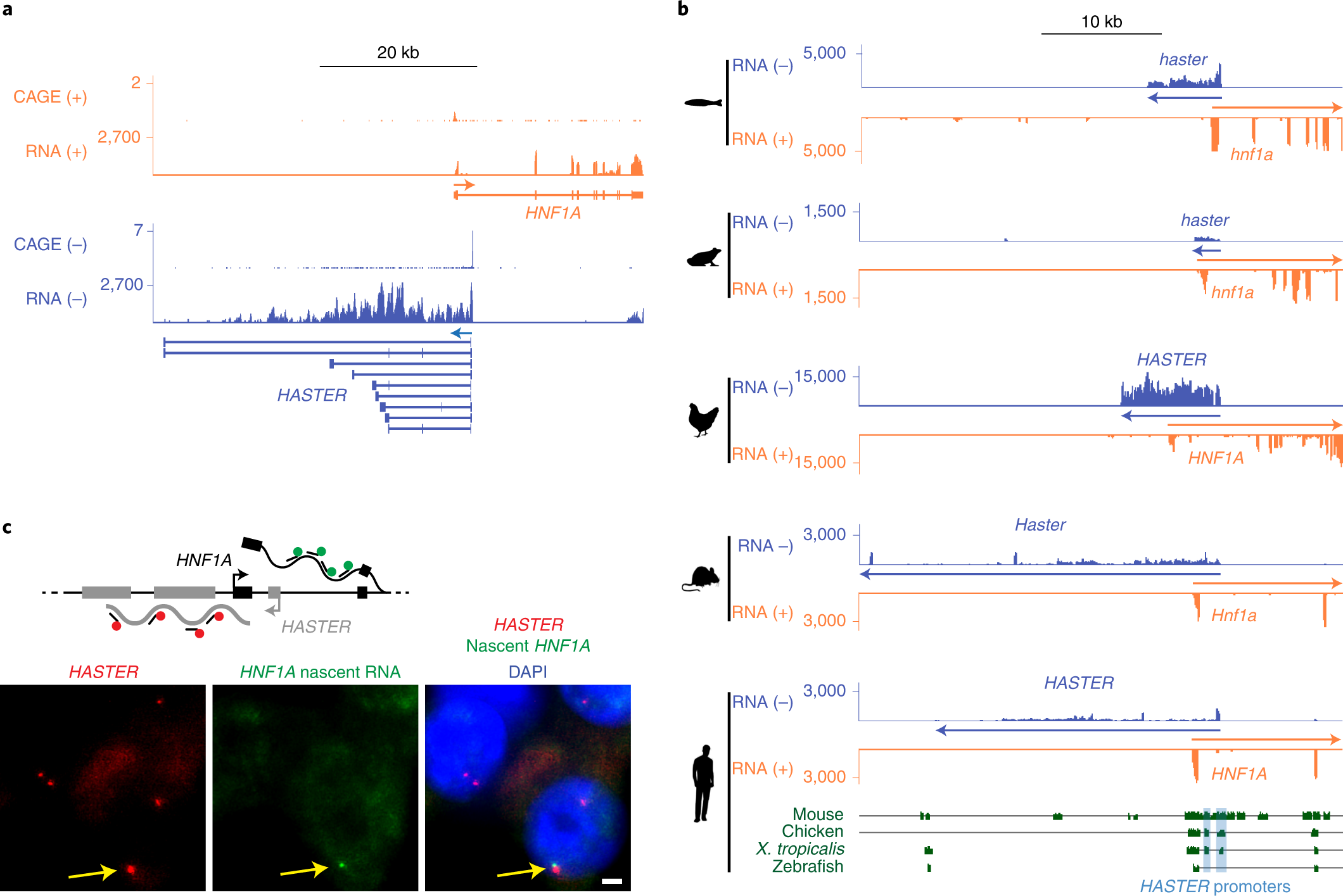
The HASTER lncRNA promoter is a cis-acting transcriptional stabilizer of HNF1A | Nature Cell Biology
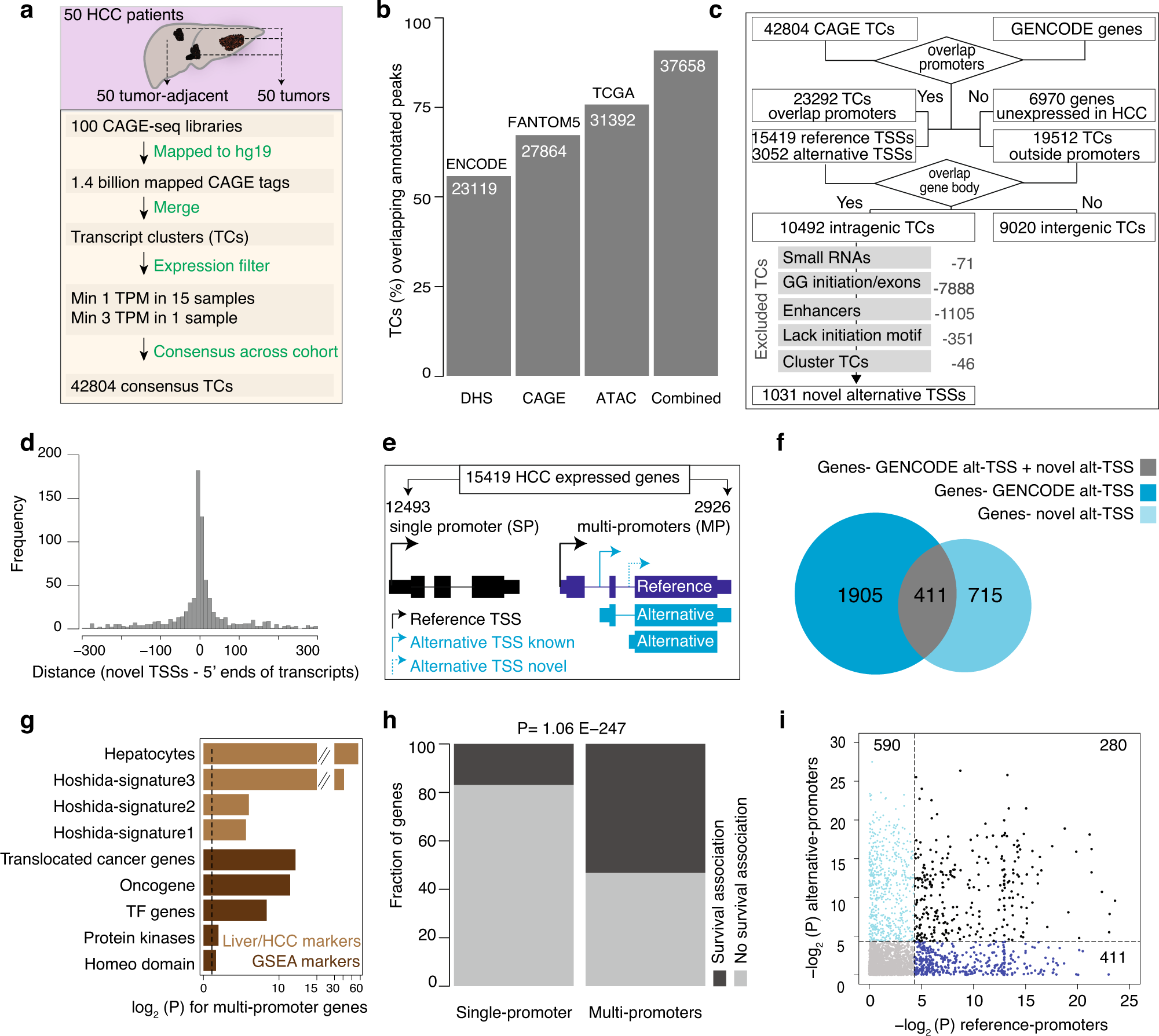
Alternative promoters in CpG depleted regions are prevalently associated with epigenetic misregulation of liver cancer transcriptomes | Nature Communications